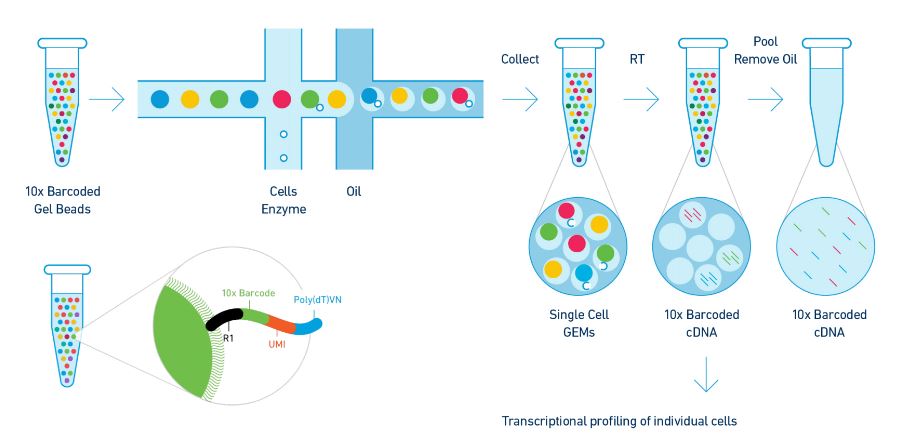
 The Chromium System provides for the massive partitioning and barcoding of single cells/nuclei using >1,000,000 unique barcodes. At the heart of the Chromium System is 10x GemCode™ Technology. Partitioning events occur on a microfluidic chip in the presence of barcoded gel beads and oil to create GEMs (Gel Bead in EMulsion). As many as 10,000 cells per sample are encapsulated in nano-liter scale GEMs. Final amplification and library construction is performed in bulk after GEMs are broken. Resulting libraries are compatible with Illumina sequencing platforms.
The Chromium System provides for the massive partitioning and barcoding of single cells/nuclei using >1,000,000 unique barcodes. At the heart of the Chromium System is 10x GemCode™ Technology. Partitioning events occur on a microfluidic chip in the presence of barcoded gel beads and oil to create GEMs (Gel Bead in EMulsion). As many as 10,000 cells per sample are encapsulated in nano-liter scale GEMs. Final amplification and library construction is performed in bulk after GEMs are broken. Resulting libraries are compatible with Illumina sequencing platforms.
Available 10x Genomic applications:
- Single Cell Digital Gene Expression
- Single Cell Immune Profiling
- Single Cell ATAC
- Visium Spatial Gene Expression
Core staff are available to assist with experimental design and to answer questions related to 10x Genomic technology. Arrangements for a meeting to discuss new or existing projects can be made by sending an email to Nathan Bivens, core director.
Assistance can also be provided by the director with grant proposals and for obtaining a letter of support from the Genomics Technology Core Facility.
The MUGTC has developed calculators to assist with the planning of sequence requirements for different applications.
Prior to sample submission and services performed, a project must be initiated with the core director and involves the following steps. Projects should be initiated at least 2 weeks prior to anticipated collection of cells/nuclei to ensure there are no issues with scheduling of an experiment. Since these reagents are very expensive and have a limited shelf life (~6 months) they are not ordered until the cell/nuclei collection is scheduled.
- Contact the core director to discuss project details. Several parameters are needed to quote a project and establish the scope of the work to be performed.
- Number of samples
- Organism and tissue for cell/nuclei source
- Number of cell/nuclei to be targeted for collection
- Number of sequence reads per cell/nuclei to be generated
- Collection date and time
- MO code for project charges
- The core director will prepare a Project Authorization Form (PAF) outlining the services. The form must be signed and returned before services are started.
- Along with the PAF, a biosafety form must also be submitted.
- Human and Animal - Use the BioSafety Form (Human and Animal)
- Plant - Use the BioSafety Form (Plant)
- A Project Information Form should be completed and submitted by email prior to sample submission.
General Considerations:
- Submit cells in Eppendorf DNA Lo Bind Tubes, 0.2 ml (Cat. No. 022431048)
- Must minimize presence of cellular aggregates, dead cells, non-cellular nucleic acids and reverse transcription inhibitors.
- On the scheduled day of submission, core staff should be notified 1 hour prior to cells/nuclei being delived. It is best practice to provide periodic updates to core staff on the day of collection.
Application Specific Requirements:
| Library Type | Quantity Required | Volume | Recommended Buffer |
Special Instructions |
|
| Single Cell 3' RNA-Seq | 0.7 - 1.2 M cells/ml | 300 µl | 1x PBS with 0.04% BSA (calcium- and magnesium-free)
Strongly recommended that buffer is purchased from following vendors.
|
|
|
| Single Cell 5' RNA-Seq and V(D)J Profiling | 0.7 - 1.2 M cells/ml | 300 µl | 1x PBS with 0.04% BSA (calcium- and magnesium-free)
Strongly recommended that buffer is purchased from following vendors.
|
|
|
| Library Type | Sequence Depth* | Sequence Type | Sequence Read | Cycle Number | |
| Single Cell 3' RNA-Seq | 20,000 reads per cell/nucleus | Paired-end, single indexing |
|
|
|
| Single Cell V(D)J Enriched | 5,000 reads per cell | Paired-end, single indexing |
|
|
|
| Single Cell 5' Gene Expression | 20,000 reads per cell | Paired-end, single indexing |
|
|
|
| Single Cell ATAC-Seq | 25,000 read pairs per nucleus | Paired-end, dual indexing |
|
|
|
| Spatial Transcriptomics | 50,000 reads per spot | Paired-end, dual indexing |
|
|
|
* minimum read count recommended by 10x Genomics; increased sequencing depth may be desired for some studies
** 1 bp shorter to accommodate NovaSeq 100 cycle kits
The 10x Genomics library preparation costs will include the cost of a Chromum chip and collection fee. Each chip will collect up to 8 samples. The per sample cost is the cost of reagents for each sample.
Example: A single cell 3' RNA-Seq project requires two collections. The first collection will involve 6 samples and the second collection 8 samples.
- 10x Genomics Chromium Chip & Collection Fee Qty. 2 $2,170
- Single Cell 3' RNA-Seq Library Prep (per sample) Qty. 14 $24,570
| Single Cell Service Items | Fee |
|
|
| NovaSeq 6000 | Fee |
|
|
The 10x Genomic Resource page provides access to product literature, app notes, and posters.
Below are links highlighting demonstrated protocols and technical notes for specific applications. A complete listing of support material can be found on the 10x Genomics Support Page.
Single Cell Gene Expression
- Single Cell Gene Expression Demonstrated Protocol Compatibility Table
- Single Cell Protocols - Cell Preparation Guide
- Isolation of Nuclei for Single Cell RNA Sequencing
- Tumor Dissociation for Single Cell RNA Sequencing
- Single Cell Suspensions from Cultured Cell Lines for Single Cell RNA Sequencing
- Guidelines for Optimal Sample Preparation
- Importance of Cell Stock Concentration for Accurate Target Cell Recovery
- Removal of Dead Cells from Single Cell Suspensions Improves Performance for 10x Genomics Single Cell Applications
- Resolving Cell Types as a Function of Read Depth and Cell Number
- Biological & Technical Variation in Single Cell Gene Expression Experiments
- Guidelines for Accurate Target Cell Counts Using 10x Genomics Single Cell Solutions
Single Cell Immuno Profiling
- Single Cell V(D)J Demonstrated Protocol Compatibility Table
- Isolation of Nuclei for Single Cell RNA Sequencing
- Tumor Dissociation for Single Cell RNA Sequencing
- Single Cell Suspensions from Cultured Cell Lines for Single Cell RNA Sequencing
- Guidelines for Optimal Sample Preparation
- Importance of Cell Stock Concentration for Accurate Target Cell Recovery
- Removal of Dead Cells from Single Cell Suspensions Improves Performance for 10x Genomics Single Cell Applications
- Resolving Cell Types as a Function of Read Depth and Cell Number
- Biological & Technical Variation in Single Cell Gene Expression Experiments Guidelines for Accurate Target Cell Counts Using 10x Genomics Single Cell Solutions
Single Cell ATAC
- Nuclei Isolation from Mouse Brain Tissue for Single Cell ATAC Sequencing
- Nuclei Isolation for Single Cell ATAC Sequencing
Spatial Gene Expression
- Visium Spatial Protocols - Tissue Preparation Guide
- Visium Spatial Gene Expression Optimized Tissues
Software
- Single Cell Software Overview
- What is Cell Ranger?
- Single Cell ATAC Software Overview
- What is Cell Ranger ATAC?
- Single Cell Immune Profiling Software Overview
- Visium Software Overview
- What is Space Ranger?
- What is Loupe Cell Browser?
- Loupe Cell Browser Gene Expression Tutorial
