The Genomics Core accommodates both single sample users and high-throughput projects. The facility utilizes a 3730xl 96-capillary DNA Analyzer with Applied Biosystems Big Dye Terminator cycle sequencing chemistry.
Read lengths up to 1100 bases per reaction are routine, providing the template is of high quality. Turn around time for submissions is < 24 hours.
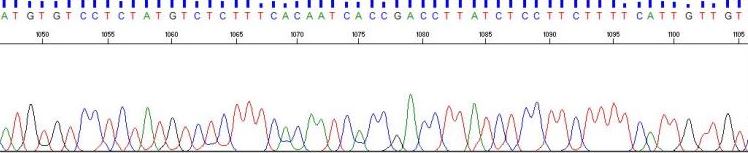
Sequence orders are placed on-line using the Genomics Core's dnaLIMS system.
First time using dnaLIMS? Use this link to create an account. Further assistance is available by contacting the Genomics Core.
Core staff are available to assist with experimental design and to answer questions related to Sanger sequencing. Arrangements for a meeting to discuss new or existing projects can be made by sending an email to Nathan Bivens, core director.
- All samples are to be submitted in a 1.5 ml tube. The rack used by our Tecan robotic system restricts us to accepting only tubes with specific dimensions. The MUGTC currently uses Fisher Scientific (Cat. #05408129) 1.5 ml MCT Graduated tubes for sample submissions. The Fisher Scientific tubes may be purchased from the Enzyme Freezer Program in a 500 tube per bag quantity.
- Total volume of the template + primer mix in the 1.5 ml tube is to be 16 µl.
- Recommended total amounts in the 16 µl volume for DNA templates and primer (strongly recommended that primer Tm be ≥ 60°C.):
- Each order receives an order number and each sample receives a requisition number. Label tubes with the last three digits of the requisition number on the top of the tube.
- Place samples in a plastic bag with the LIMS order number and submitting individual's name clearly written on the outside of the bag. (Plastic bags and permanent marker are provided on a cart outside of room 215).
- Place samples in the white refrigeration unit located in the hallway directly across from room 211. A box for samples will be on the left side of the unit on the top shelf.
| DNA Type |
DNA Length |
DNA total mass (nanograms) |
Primer Total (picomoles) |
||
| Plasmid DNA(dsDNA) | < 5 kb | 500 - 750 ng | 20 pmoles | ||
| 5 - 10 kb | 750 - 1000 ng | 20 pmoles | |||
| > 10 kb | < 1200 ng | 20 pmoles | |||
| Single Stranded DNA | 275 - 500 ng | 20 pmoles | |||
| Purified PCR Product | < 500 bp | 20 -50 ng | 20 pmoles | ||
| 500 - 1000 bp | 50 -100 ng | 20 pmoles | |||
| 1000 - 2000 bp | 100 -250 ng | 20 pmoles | |||
| 2000 - 5000 bp | 250 - 500 ng | 20 pmoles | |||
| 5000 bp - 10 kb | 750 - 1000 ng | 20 pmoles | |||
|
Large DNA |
2000 ng | 20 pmoles | |||
|
For example: Order #15759 has two sample; Requisition #289573 and Requisition #289574. Each tube is labeled with the last three digits of the requisition #. Requisition #289573 Requisition #289574 |
 |
Considerations for 96-well Plate Submission:
- Samples are to be submitted in a 96-well plate obtained from the Genomics Core. Plates containing less than 96 samples should be loaded in a vertical fashion (A1, B1, C1, D1, E1, F1, G1, H1, A2, B2, etc.)
- Total volume of the template + primer mix in each well of the plate is to be 6 µl.
- Add 5 - 10 pmol of a single primer to each reaction.
- The 3730 is sensitive to salts and other contaminants. Care in template preparation is essential to quality data and instrument maintenance.
| Submission Type | Fee |
|
|
MUGTC Troubleshooting Guide - This on-line guide provides examples of common sequencing issues and recommendations. Staff are available for consultation or to review data.
QIAGEN Guide to Template Purification and DNA Sequencing - This guide provides examples of common sequencing issues and recommendations. Staff are available for consultation or to review data.
Software
The following software is a list of known freeware tracefile viewers. The MUGTC doesn't monitor updates of these programs nor make any claims to functionality. Please read the documentation to ensure compatibility with your computer operating system and to understand the features offered by each software package.
| Software | Windows | Mac | Linux | Website Link | |
| Chromas 2.6.6 | Windows XP, Vista, 7, 8, 10, 11 | - | - |  |
|
| 4Peaks | - | OS X 10.7 + | - |  |
|
| SnapGene | Windows 7, 10 | OS 10.10 + | Linux 7.2 |  |
|
| UGENE | Windows XP, Vista, 7, 8 | OS X | Linux |  |
|
 How can contaminants be avoided with the use of spin columns in the isolation method?
How can contaminants be avoided with the use of spin columns in the isolation method?
 Why does the Genomics Core require the use of a specific tube for Sanger sequencing submissions?
Why does the Genomics Core require the use of a specific tube for Sanger sequencing submissions?
 Why does the Genomics Core require a volume of 16 µl for sequencing submissions?
Why does the Genomics Core require a volume of 16 µl for sequencing submissions?
 Why is it that only a portion of my data from a single order is available? When will I receive the remaining data?
Why is it that only a portion of my data from a single order is available? When will I receive the remaining data?
 How can I distinguish results that may be background noise from results with signal intensity?
How can I distinguish results that may be background noise from results with signal intensity?
 Several samples have been submitted but some of the samples in the order will not be taken to the core. Can I delete the individual samples from the order?
Several samples have been submitted but some of the samples in the order will not be taken to the core. Can I delete the individual samples from the order?
 How much DMSO should I add to the 16ul submission volume to acheive a 5% concentration in the reaction?
How much DMSO should I add to the 16ul submission volume to acheive a 5% concentration in the reaction?
 Is there cause for secondary structure to occur at the cloning site when using pCR™II and pCR™II vector?
Is there cause for secondary structure to occur at the cloning site when using pCR™II and pCR™II vector?
